Welcome!
Email: jjian [at] scripps [dot] edu
Hi there! I’m Jinglin Jian (简靖琳), a PhD student at Scripps Research by the beautiful ocean 🏖️ at San Diego, CA. I’m deeply grateful to be supported by the Kellogg Fellowship, a three-year endowed award generously funded by the Kellogg family and The ALSAM Foundation.
I received my master’s degree from the School of Information Sciences at the University of Illinois at Urbana-Champaign, where I had the opportunity to work closely with Professor Ge Liu and Professor Qingyun Wang.
I’m also the founder of the AI × Science Club, a student-led local community exploring how AI is reshaping scientific discovery. We’re still growing and building things out, so if this sounds interesting to you, come join us on Slack — we’d love to have you!
News
- Feb 2026: Founded the AI × Science Club at Scripps Research. We’re still building, if you’re interested, come join us on Slack!
- Feb 2026: Rotating in Bryan Briney’s lab, working on antibody language models.
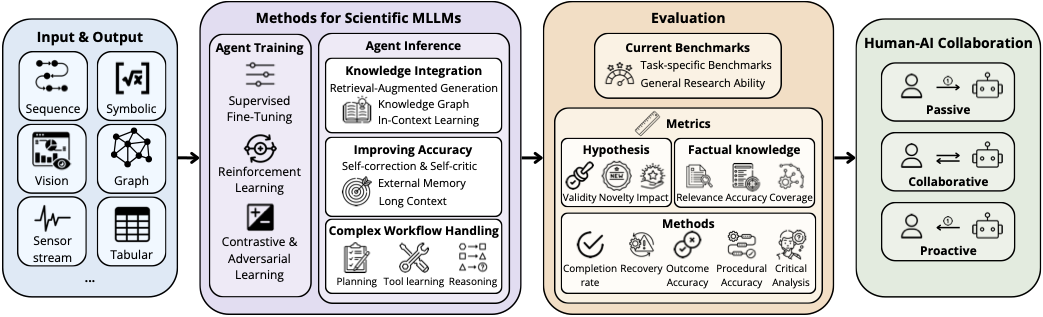
- Dec 2025: Survey paper “Exploring Agentic Multimodal LLMs for AI Scientists” published on TechRxiv.
- Oct 2025: Selected for the three-year Kellogg Fellowship at Scripps Research. See here.
- Aug 2025: Started PhD study at Scripps Research, rotating in Stefano Forli’s lab.
- May 2025: Graduated from UIUC with MS in Information Sciences.
- Nov 2024: Two papers accepted to IEEE BigData 2024, including one as first author.
Research Interests
I am mainly interested in pretraining models on molecular dynamics data (3D with time series) and protein sequence data (3D static data).
Publications and Conferences
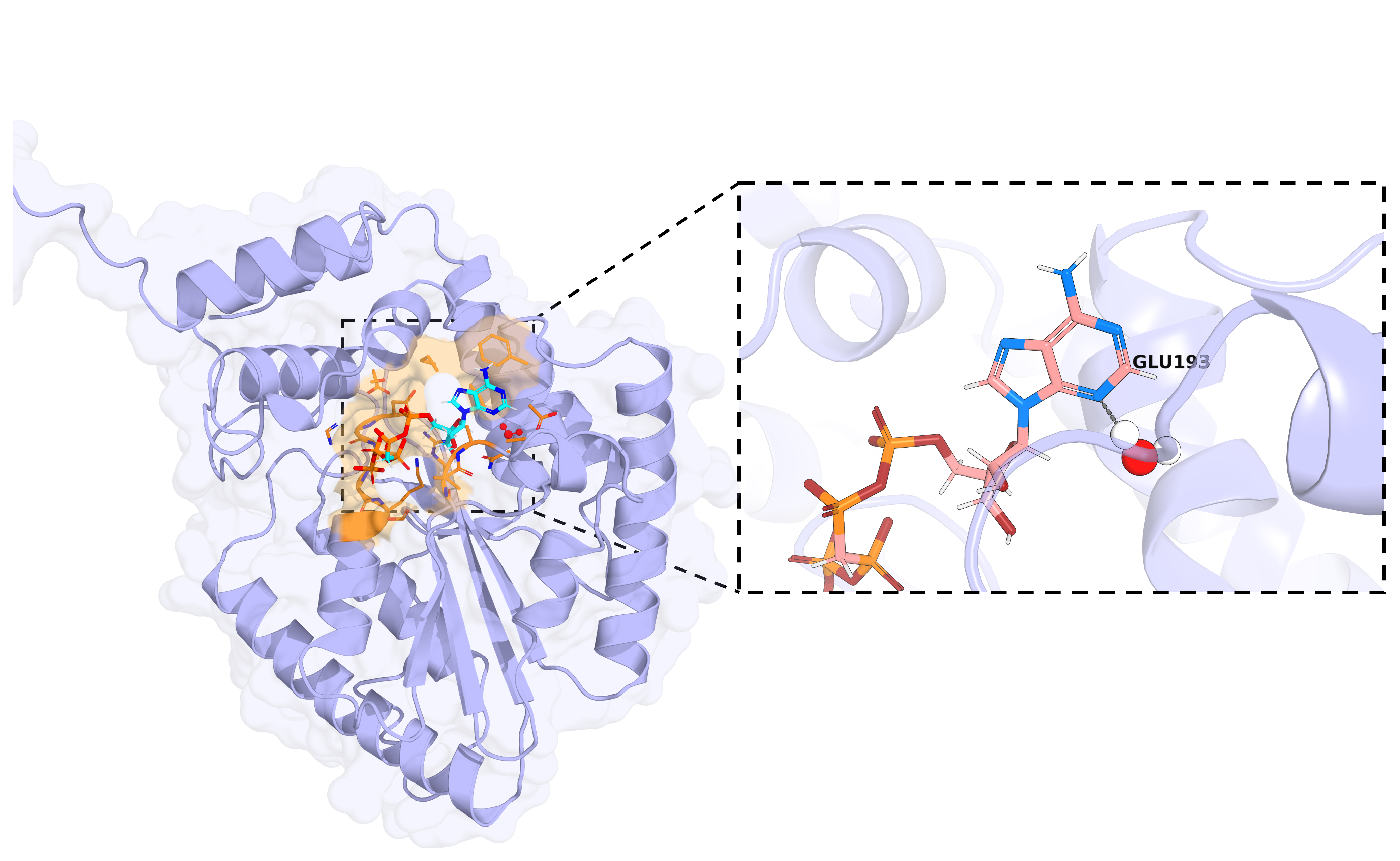
Allosterically Inhibiting Pseudomonas aeruginosa's Polyphosphate Kinase 2A by Disrupting Its Oligomerization
Constanza Torres-Paris, Madeline G. Ammend, Joseph Agha, Jinglin Jian, Matthew Holcomb, Stefano Forli, Lisa R. Racki
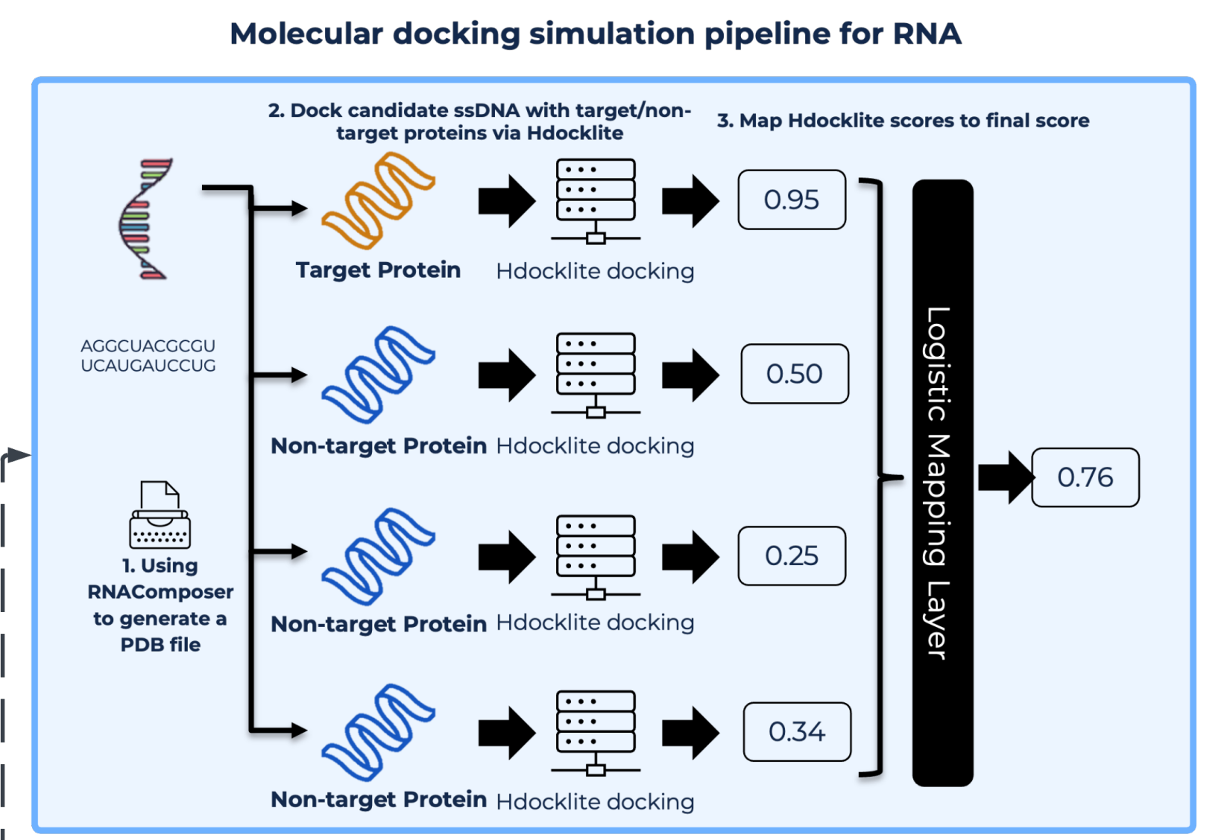
Performed virtual screening of 2.4 million compounds against Ppk2A mutant proteins using AutoDock-GPU and identified preliminary hit candidates.
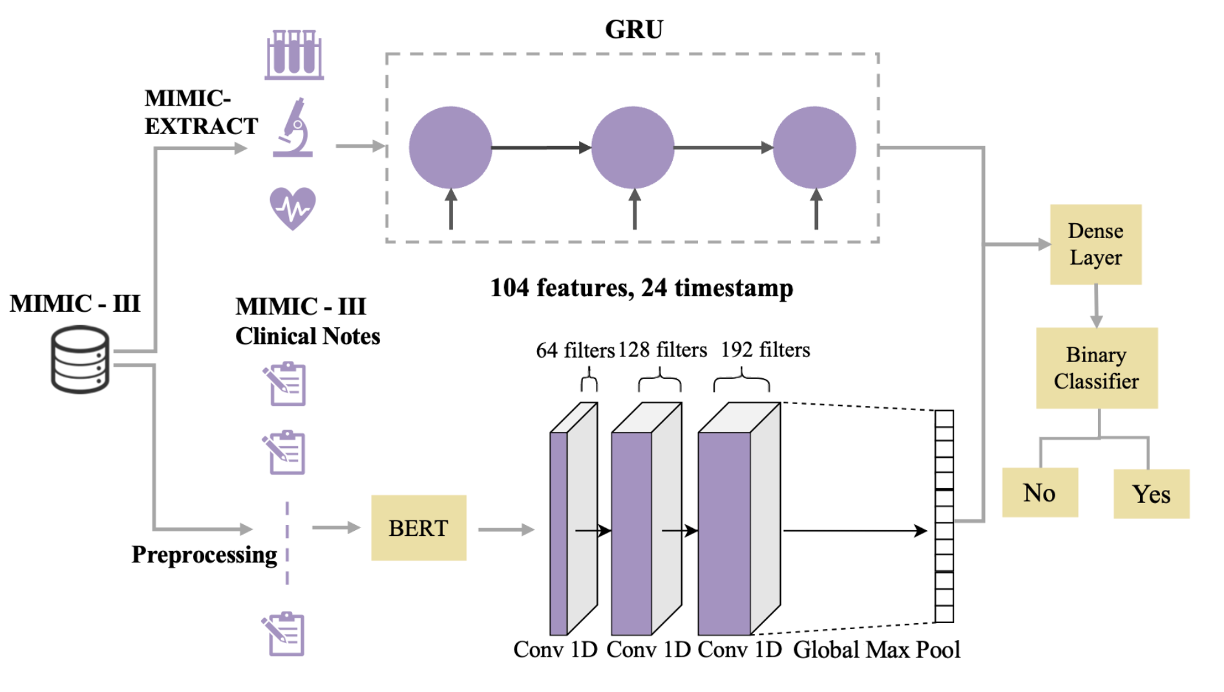
Patient Outcome Predictions via A Multimodal Language Model for Electronic Health Records
Zihan Li, Jinglin Jian, Chundian Li, Jinxia Yao, Jin Chen, Yang Zhang
Early prediction of mortality risk and hospital length of stay is critical. We propose a multimodal framework that integrates full-text clinical note embeddings and time-stamped physiological data to jointly model patient outcomes.
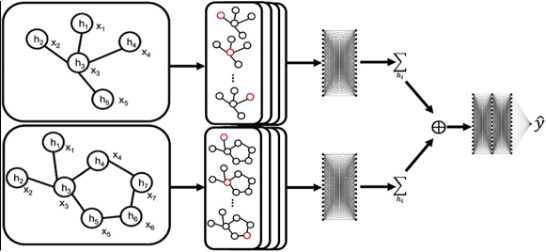
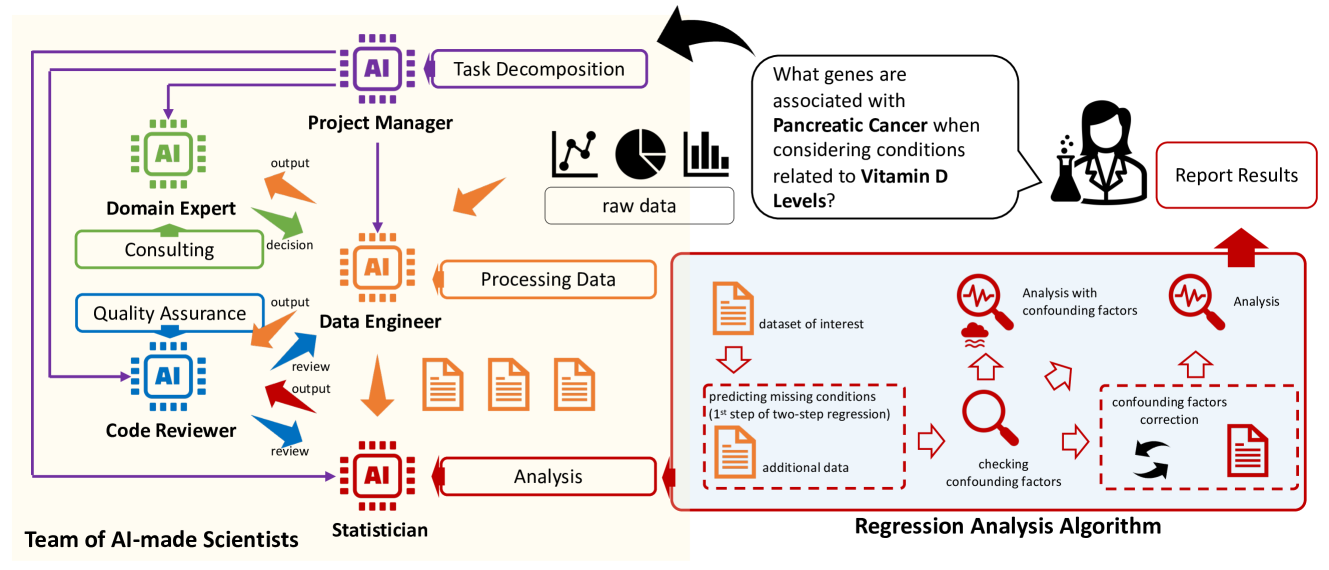
Toward a Team of AI-made Scientists for Scientific Discovery from Gene Expression Data
Haoyang Liu, Yijiang Li, Jinglin Jian, Yuxuan Cheng, Jianrong Lu, Shuyi Guo, Jinglei Zhu, Mianchen Zhang, Miantong Zhang, Haohan Wang
ML can discover disease-predictive genes from gene expression data. We introduced the Team of AI-made Scientists (TAIS), a LLM-based framework for automatic streamlining ML analysis. TAIS consists of simulated roles, including a project manager, data engineer, and domain expert.
Selected Projects
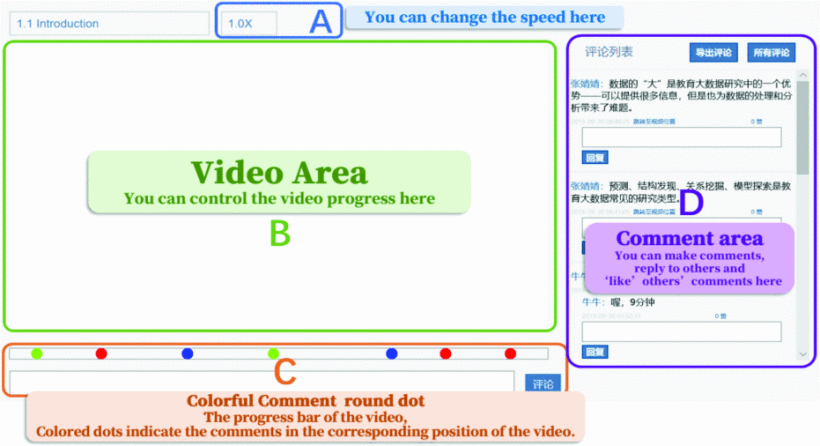
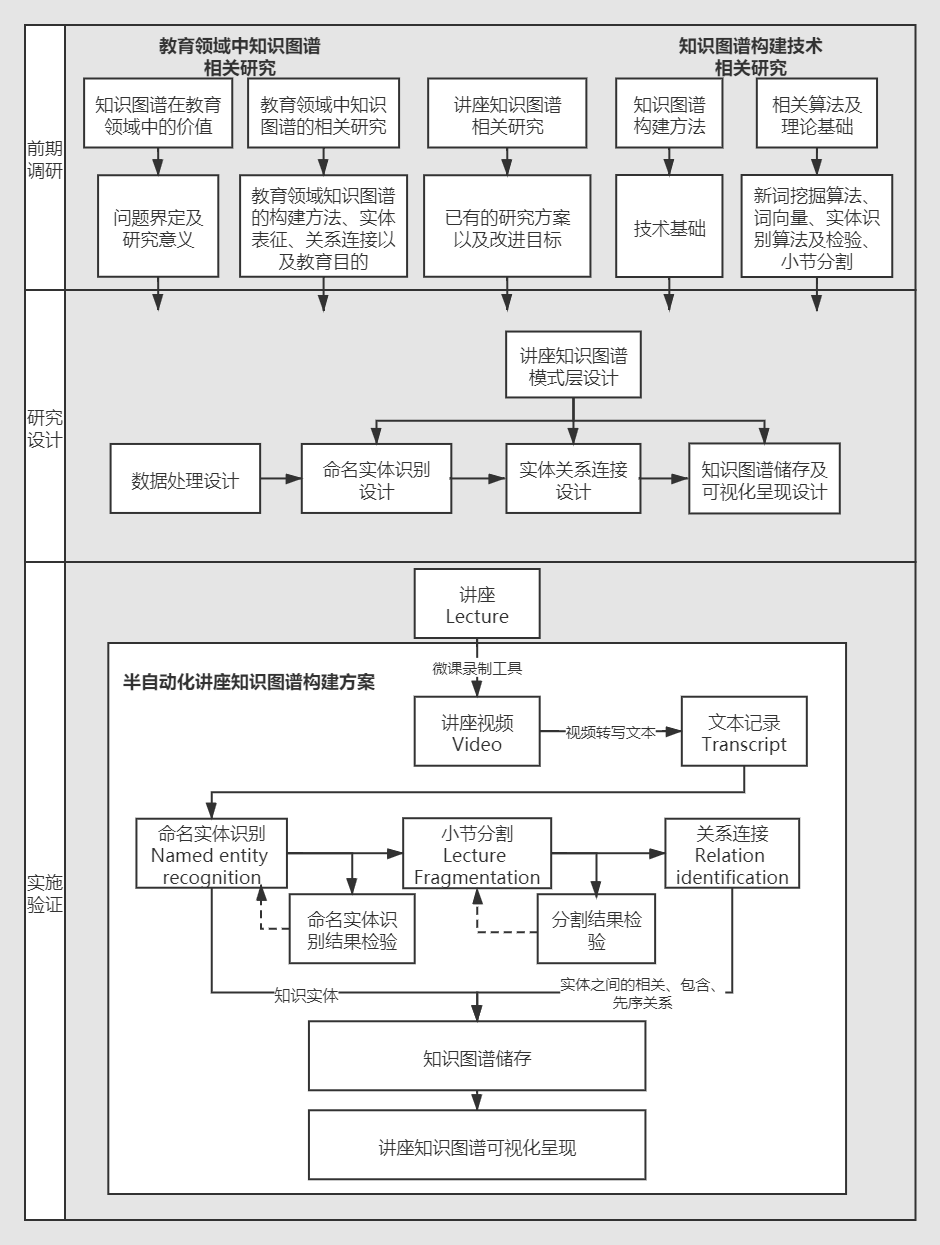
The Impact of Productive Failure on Learning Performance and Cognitive Load: Using Hypervideo to Facilitate Online Interactions
Xiaojie Niu, Jingjing Zhang, Kate M. Xu, Xuan Wang
Productive failure is an instructional approach that uses students’ cognitive conflicts to enhance their learning. This experimental study investigated the effect of productive failure in a hypervideo environment.